EpiGraph Interactive Project Page · Paper: arXiv:2605.09505
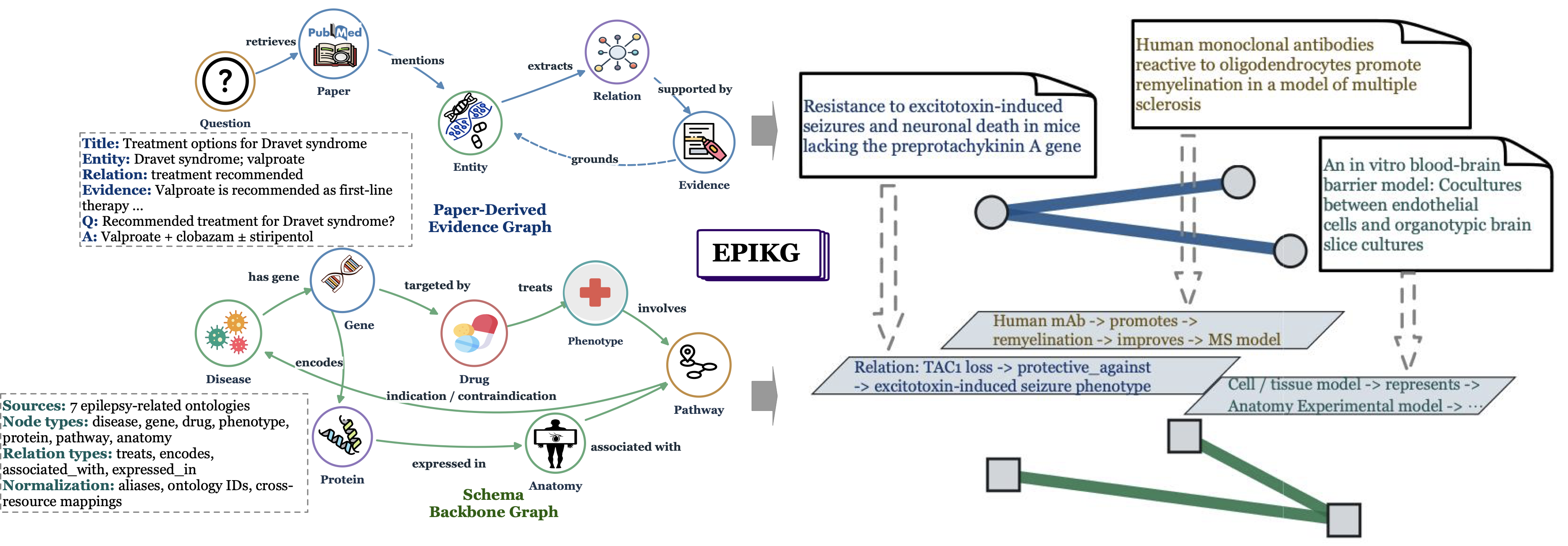
48,166 Papers · 24,324 Entities · 32,009 Triplets · 5 Evidence-Intensive Epilepsy Reasoning Tasks
How to Cite · News · Why EpiGraph · Key Features · Quick Start · Tasks · Metrics
EpiGraph Interactive Project Page · Paper: arXiv:2605.09505
How to Cite · News · Why EpiGraph · Key Features · Quick Start · Tasks · Metrics
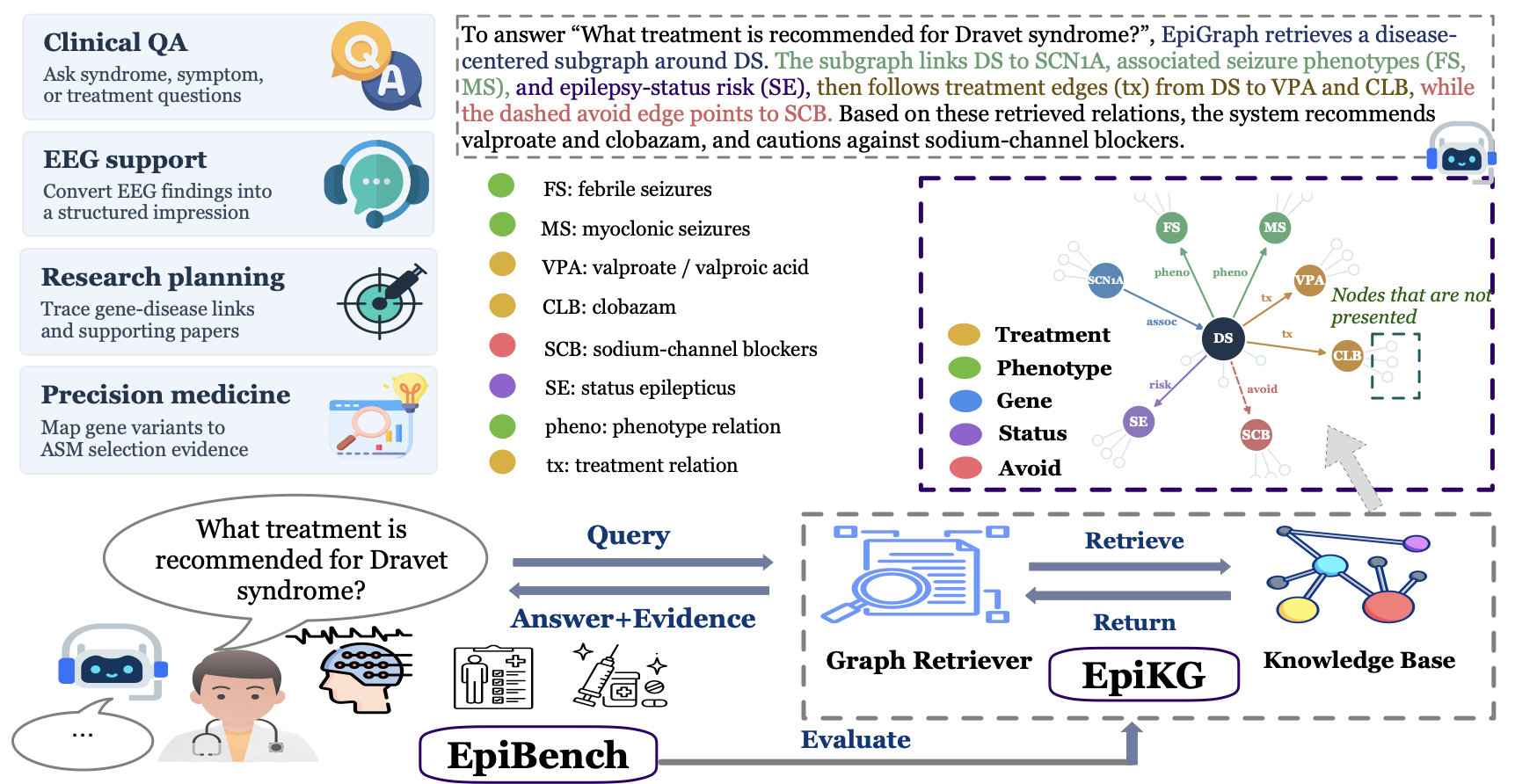
Interactive knowledge graphExplore a compact EpiGraph subgraph directly in the browser. Search nodes, inspect evidence paths, and view relation metadata used by Graph-RAG. |
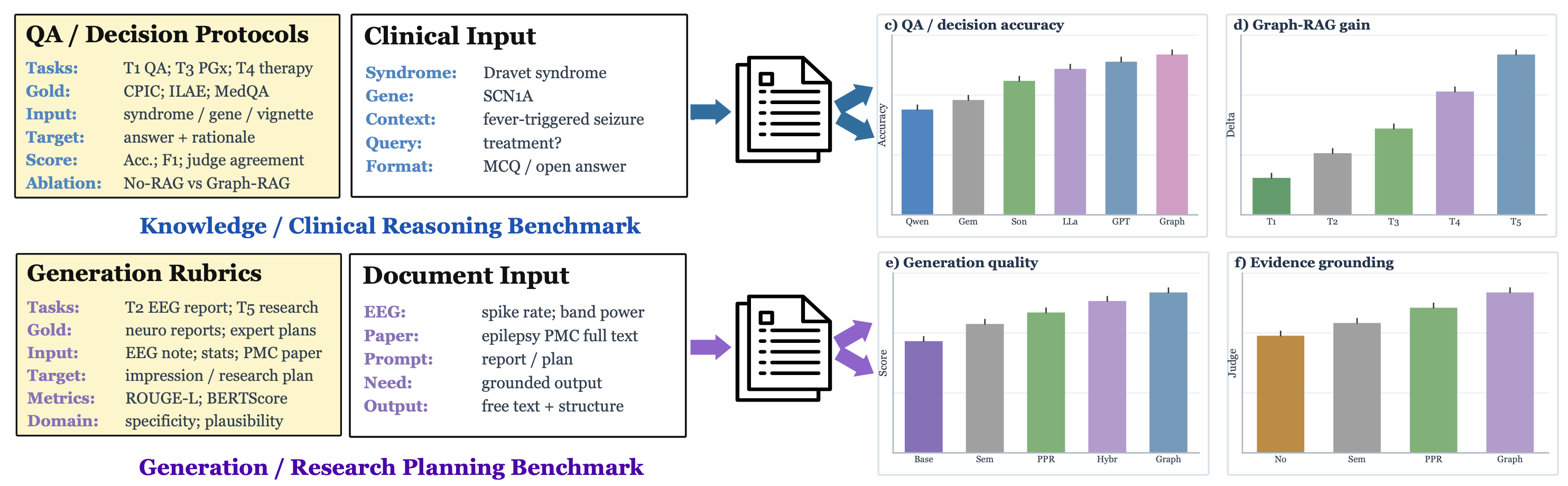
Plug-and-play evaluationRun the same task scripts with your own model, retriever, prompts, or local data exports. EpiBench is designed for fast model testing and fair ablation. |
Five clinically grounded tasksEvaluate models on epilepsy diagnosis, EEG impression generation, biomarker-driven medication selection, treatment recommendation, and deep research planning. |
Private-data-aware releaseThe Harvard EEG task is supported through a local schema adapter, so the evaluation logic is reproducible without redistributing restricted data. |